Generate Posterior Simulations
Source:vignettes/02-posterior-simulation.Rmd
02-posterior-simulation.RmdGenerate Posterior Simulations
This vignette shows how to generate synthetic climate data from fitted posteriors, then fit BFACT to evaluate model sensitivity.
Workflow:
- Extract posterior samples from a model fitted to real data (e.g., )
- Utilize parameters from these posterior samples to generate synthetic climate trajectories
- Fit BFACT with multiple values (1-20) to the synthetic data
Step 1: Fit Real Data and Extract Posterior Samples
Start by fitting BFACT to the real New York temperature data, then extract posterior draws. Here .
library(BFACT)
library(ncdf4)
library(dplyr)
# Load real climate data (same as vignette 01)
nc_file <- system.file("data", "NewYork_temperature_anomalies_JJA_all_models_1961-1990baseline.nc", package = "BFACT")
nc <- nc_open(nc_file)
temp_data <- ncvar_get(nc, "temperature")
years <- ncvar_get(nc, "year")
models <- ncvar_get(nc, "model")
temp_avg <- apply(temp_data, c(3, 4, 5), mean, na.rm = TRUE)
# Use SSP 5-8.5 (highest warming scenario)
Y_real <- temp_avg[, 3, ]
# Add model names as column names - this is necessary for outlier detection and later analysis
colnames(Y_real) <- models
# Remove outlier models
outliers <- BFACT::detect_outliers(
data = Y_real,
start_index = 1,
end_index = nrow(Y_real),
threshold_fn = BFACT::mean_sd_threshold,
threshold_dial = 2.5
)
if (length(outliers) > 0) {
cat("Removed outlier models:", paste(outliers, collapse = ", "), "\n")
Y_real <- Y_real[, !colnames(Y_real) %in% outliers, drop = FALSE]
}## Removed outlier models: CIESM, UKESM1-0-LL
hadcrut_file <- system.file("data", "hadcrut5_annual.rds", package = "BFACT")
z_real <- readRDS(hadcrut_file)
nc_close(nc)
# Fit BFACT to real data with H0=2
cat("Fitting BFACT to real data (H0=2)...\n")## Fitting BFACT to real data (H0=2)...
fit_real <- BFACT(
Y = Y_real,
z = z_real,
T = 251,
T1 = 1,
T2 = 173,
H = 2,
iseed = 123,
J = 6,
nsim = 100
)Step 2: Generate Synthetic Data from Posterior Samples
# Generate synthetic data from this posterior draw
sim_posterior <- simulate_from_posterior(
fit = fit_real,
years_start = 1850,
years_end = 2100,
T1 = 1,
T2 = 173,
J = 6,
OBS_COL = 1,
seed = 123
)
cat("Real data fit complete. Generated synthetic observations from H0=2 posterior.\n")## Real data fit complete. Generated synthetic observations from H0=2 posterior.
# Plot simulated data with true trend
p_sim <- plot_data_with_posterior(
Y = sim_posterior$Y,
z = sim_posterior$z,
true_trend = sim_posterior$mu[, 1],
title = "Synthetic Data: Generated from H0=2 Posterior",
years = sim_posterior$years,
obs_years = sim_posterior$T1:sim_posterior$T2
)
p_sim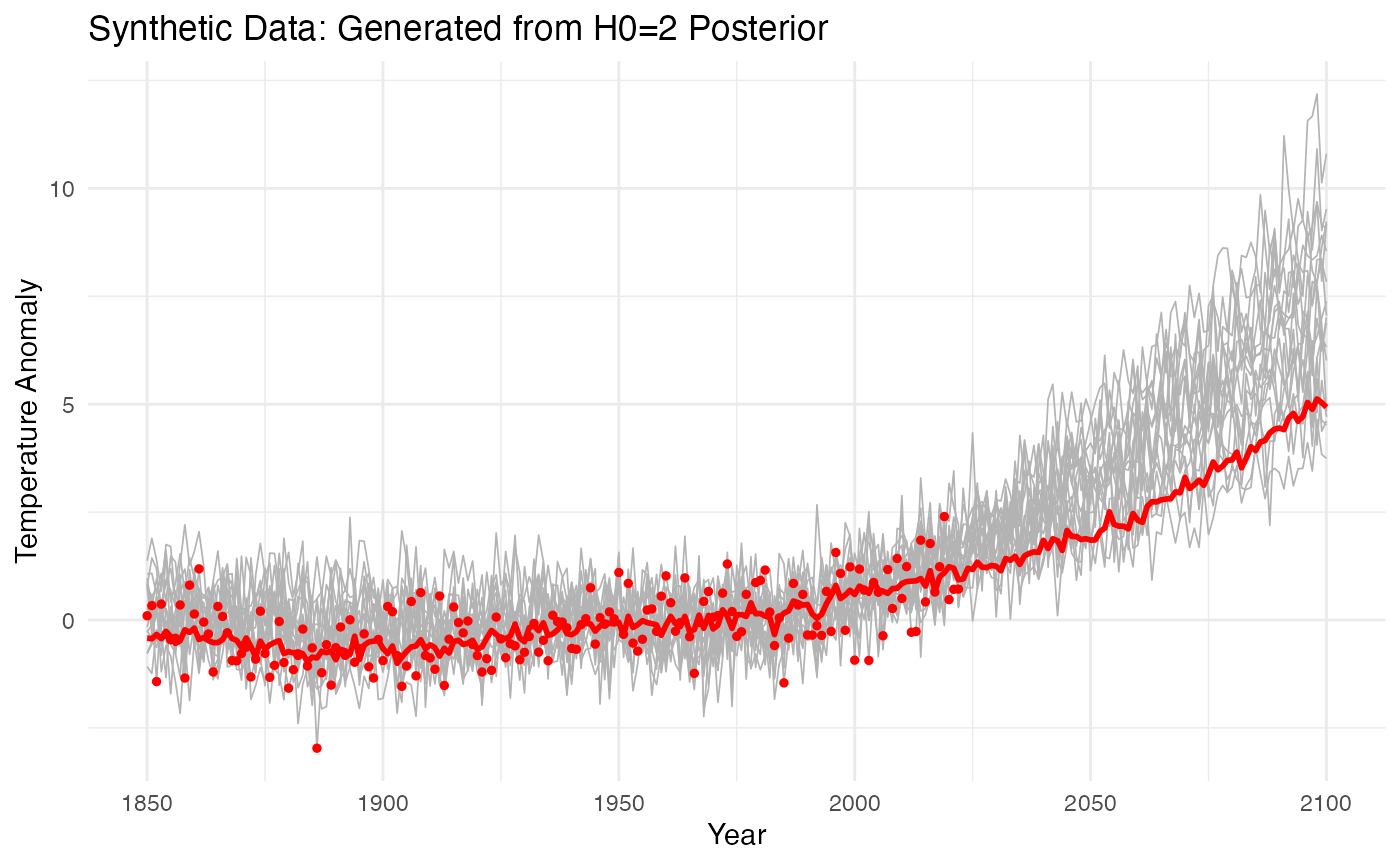
Step 3: Fit Multiple H Values to Synthetic Data
Now fit BFACT with to evaluate model sensitivity:
# Fit H=2 to H=5 to the synthetic data
H_fit_range <- 2:5
fits_posterior <- list()
for (H in H_fit_range) {
cat(sprintf("\nFitting H=%d to synthetic data from H=2 posterior...\n", H))
fit <- BFACT(
Y = sim_posterior$Y,
z = sim_posterior$z,
T = 251,
T1 = 1,
T2 = 173,
H = H,
iseed = 123 + H,
J = 6,
nsim = 100
)
fits_posterior[[as.character(H)]] <- fit
}##
## Fitting H=2 to synthetic data from H=2 posterior...
##
## Fitting H=3 to synthetic data from H=2 posterior...
##
## Fitting H=4 to synthetic data from H=2 posterior...
##
## Fitting H=5 to synthetic data from H=2 posterior...
cat("Completed fits for H=2:5\n")## Completed fits for H=2:5Plotting Posterior Fits
library(patchwork)
# Generate posterior fit plots for each H
plots_H <- list()
for (H in H_fit_range) {
fit <- fits_posterior[[as.character(H)]]
plots_H[[as.character(H)]] <- plot_data_with_posterior(
Y = sim_posterior$Y,
z = sim_posterior$z,
posterior_samples = sample_posterior(fit),
title = sprintf("H = %d", H),
years = sim_posterior$years,
obs_years = sim_posterior$T1:sim_posterior$T2
)
# Only keep y-axis label on the middle plot
if (H != 3) {
plots_H[[as.character(H)]] <- plots_H[[as.character(H)]] + ggplot2::labs(y = NULL)
}
}
# Combine plots in grid
grid_plot <- plots_H[[1]] / plots_H[[2]] / plots_H[[3]] / plots_H[[4]]
print(grid_plot)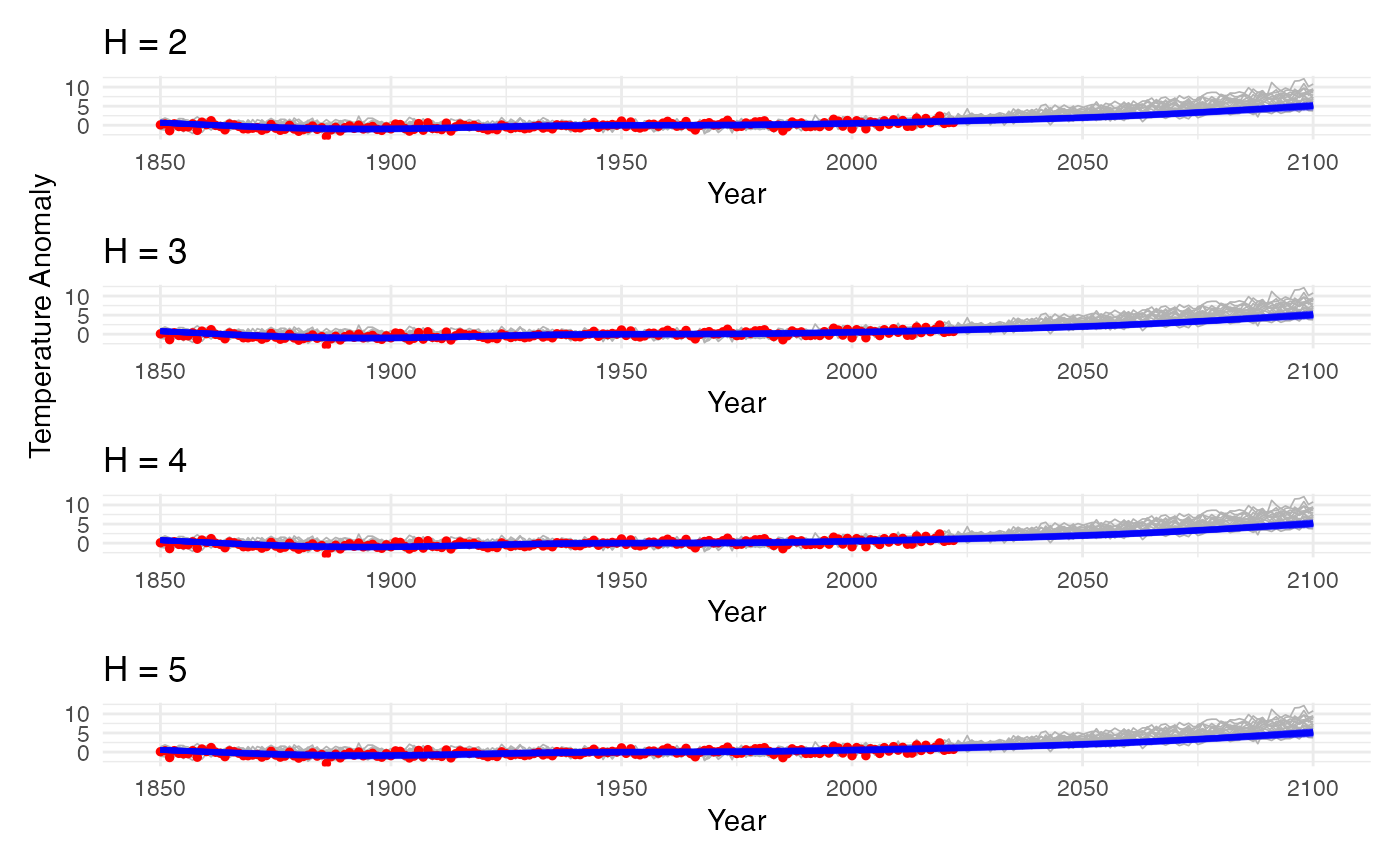
Step 4: Model Sensitivity to H
Compute RMSE for each value and assess.
out <- consolidate_results(fits_posterior, sim_posterior)
cat("\nModel comparison (RMSE on synthetic data):\n")##
## Model comparison (RMSE on synthetic data):
print(out$model_comparison)## # A tibble: 4 × 4
## H RMSE mean_post_var max_post_var
## <int> <dbl> <dbl> <dbl>
## 1 2 0.239 0.0102 0.0530
## 2 3 0.260 0.00960 0.0516
## 3 4 0.258 0.0102 0.0483
## 4 5 0.223 0.0126 0.0520